Clustering Methods
Discover hidden groups and patterns in unlabeled data using proven clustering algorithms
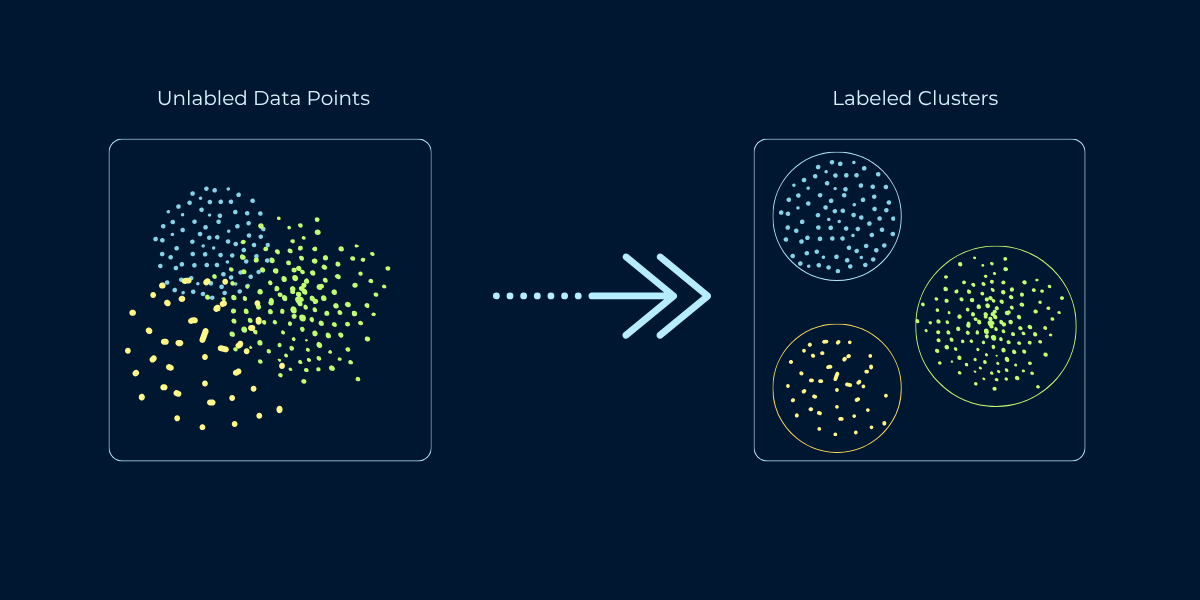
You've mastered supervised learning, where algorithms learn from labeled examples. Now explore the opposite: unsupervised learning, where algorithms discover patterns without any labels. Clustering is the most fundamental unsupervised learning task - finding natural groups in data when you don't know the groups in advance. When Netflix groups similar viewers, when Amazon segments customers by shopping behavior, when researchers discover new cancer subtypes from genetic data - that's clustering revealing hidden structure in unlabeled data.
What Makes This Tutorial Different
Clustering is fundamental because it's how we make sense of the world. Every concept you have - from 'chair' to 'democracy' - is really a cluster of similar instances. When machines can cluster as well as humans do, they'll truly understand. The challenge isn't just finding clusters, it's finding the right clusters at the right level of abstraction.

What Is Clustering?
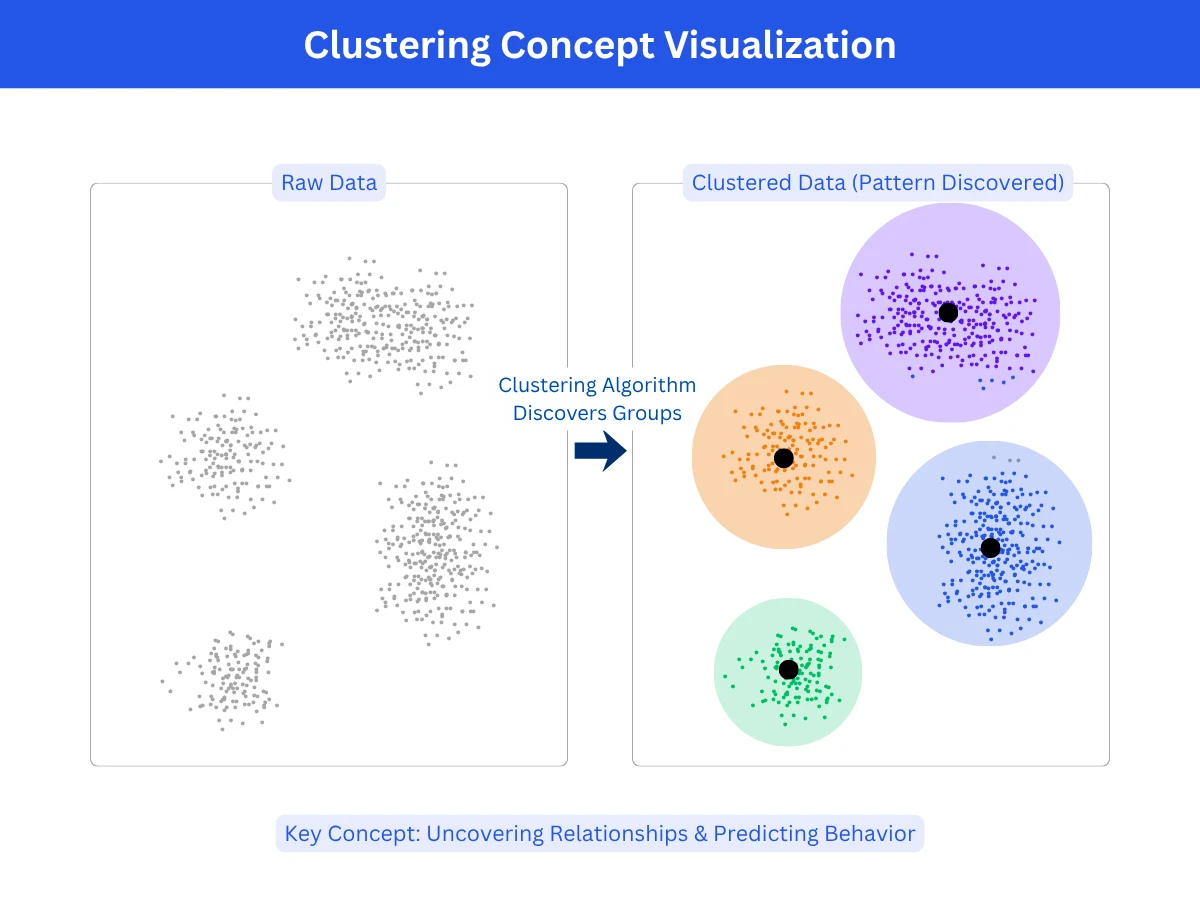
Clustering is the task of grouping similar data points together without predefined categories or labels. The algorithm examines features of your data and discovers natural groupings based on similarity. Points within a cluster should be similar to each other and different from points in other clusters. Unlike classification, where you predict which predefined category a data point belongs to, clustering discovers the categories themselves from the data's structure.
Think of organizing a photo library. Classification would be: given labeled categories (family, vacation, work), assign each new photo to a category. Clustering would be: look at all photos and group similar ones together, discovering natural categories like "beach photos," "indoor group shots," "sunset landscapes" without being told these categories exist. The algorithm finds structure you didn't know was there.
Types of Clustering Problems
- Partitional clustering: Divide data into non-overlapping groups where each point belongs to exactly one cluster. Example: k-means assigns every customer to one segment (budget, mid-tier, premium).
- Hierarchical clustering: Build a tree of nested clusters, allowing different granularities. Example: biology taxonomy (species - genus - family - order) where clusters nest within larger clusters.
- Density-based clustering: Find regions of high density separated by low-density regions. Points in sparse areas become outliers. Example: detecting fraud patterns - normal transactions cluster densely, fraudulent ones are sparse outliers.
- Model-based clustering: Assumes each cluster has a natural shape and finds groups by fitting those shapes to the data. Example: if you plot customer spending, you might see two humps - frequent small purchases and occasional large ones. The algorithm fits a curve to each hump to define the groups, so a customer near the overlap can softly belong to both rather than being forced into one.
Clustering vs Classification: Key Differences
Clustering and classification both group data, but they approach the problem from opposite directions. Understanding this distinction is crucial for choosing the right technique for your problem.
| Aspect | Classification (Supervised) | Clustering (Unsupervised) |
|---|---|---|
| Learning Type | Supervised - requires labeled training data | Unsupervised - no labels needed |
| Categories | Predefined - you specify categories in advance | Discovered - algorithm finds categories from data |
| Goal | Predict which known category new data belongs to | Discover unknown groups and patterns in data |
| Training Data | Examples with known labels (spam/not spam) | Just features, no labels |
| Number of Groups | Fixed - determined by training labels | Variable - you choose or algorithm decides |
| Evaluation | Accuracy against known labels | Cluster quality metrics (cohesion, separation) |
| Example | Classify email as spam using labeled training emails | Group customers by behavior without predefined segments |
Quick Decision Rule
Common Clustering Algorithms
Different clustering algorithms make different assumptions about data structure. Understanding each algorithm's approach helps you choose the right one for your specific data characteristics and goals.
1. K-Means Clustering
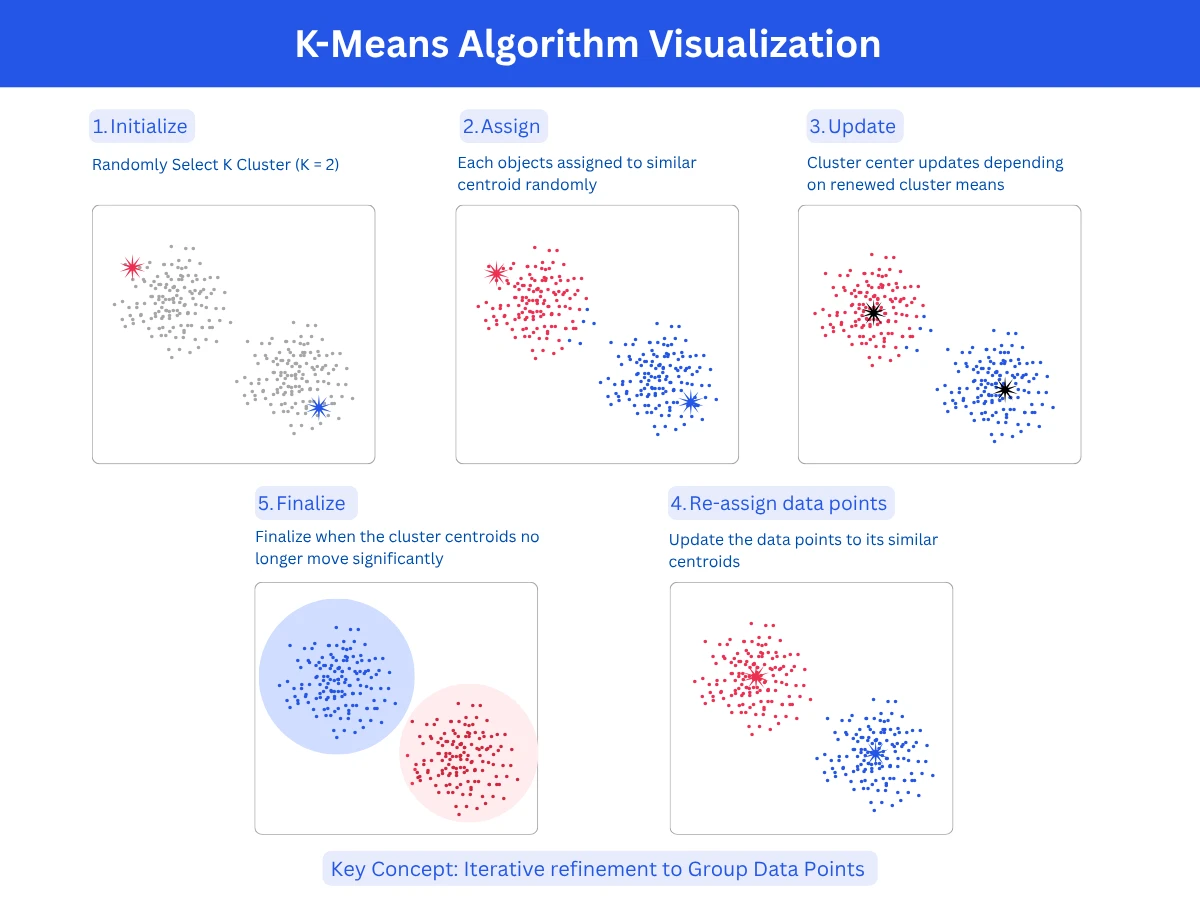
K-means is the most popular clustering algorithm due to its simplicity and speed. You specify K (the number of clusters), and the algorithm assigns each point to the nearest cluster center (CentroidCentroidThe center point of a cluster, calculated as the average position of all points in that cluster. The algorithm moves centroids around until they settle in the best position.Example:Like finding the geographic center of a city by averaging all the coordinates of its buildings.), then iteratively refines centroids until ConvergenceConvergenceWhen a model stops improving during training because it has found the best possible solution. The learning process has "converged" to a stable answer.Example:Like adjusting a recipe - after 5 tweaks it tastes perfect, so you stop adjusting.. It works by minimizing the distance from points to their assigned centroid. The algorithm is fast, scales well to large datasets, and produces Spherical ClustersSpherical ClustersClusters that are roughly round and equal-sized, like bubbles. K-means assumes this shape, so it struggles when clusters are elongated, irregular, or very different in size.Example:A scatter plot of customer age vs income forming three circular blobs - those are spherical clusters., evenly-sized clusters.
- How it works: Initialize K random centroids, assign each point to nearest centroid, recalculate centroids as cluster means, repeat until centroids stabilize
- Strengths: Very fast (scales to millions of points), simple to understand and implement, guaranteed to converge, works well for spherical clusters
- Limitations: Must specify K in advance, assumes spherical equal-sized clusters, sensitive to initial centroid placement, struggles with irregular shapes
- Best for: Large datasets, spherical well-separated clusters, when K is known or can be estimated
- Common uses: Customer segmentation, image compression, document clustering, feature engineering
1
from sklearn.cluster import KMeans
2
from sklearn.datasets import make_blobs
3
import matplotlib.pyplot as plt
4
import numpy as np
5
6
# Generate synthetic data with 3 natural clusters
7
X, y_true = make_blobs(n_samples=300, centers=3, cluster_std=0.6, random_state=42)
8
9
# Apply K-Means clustering
10
kmeans = KMeans(n_clusters=3, random_state=42, n_init=10)
11
kmeans.fit(X)
12
13
# Get cluster assignments and centroids
14
labels = kmeans.labels_
15
centroids = kmeans.cluster_centers_
16
17
print(f"Cluster assignments (first 10 points): {labels[:10]}")
18
# Output: [0 0 2 2 0 0 1 1 2 2]
19
20
print(f"\nCluster centroids:")
21
for i, centroid in enumerate(centroids):
22
print(f" Cluster {i}: [{centroid[0]:.2f}, {centroid[1]:.2f}]")
23
# Output:
24
# Cluster 0: [-8.94, -8.48]
25
# Cluster 1: [4.02, -8.08]
26
# Cluster 2: [1.83, -0.15]
27
28
# Evaluate clustering quality
29
inertia = kmeans.inertia_ # Sum of squared distances to nearest centroid
30
print(f"\nInertia (lower is better): {inertia:.2f}") # Output: 327.45
31
32
# Predict cluster for new point
33
new_point = [[0, 0]]
34
predicted_cluster = kmeans.predict(new_point)
35
print(f"\nNew point [0, 0] belongs to cluster: {predicted_cluster[0]}")
2. Hierarchical Clustering
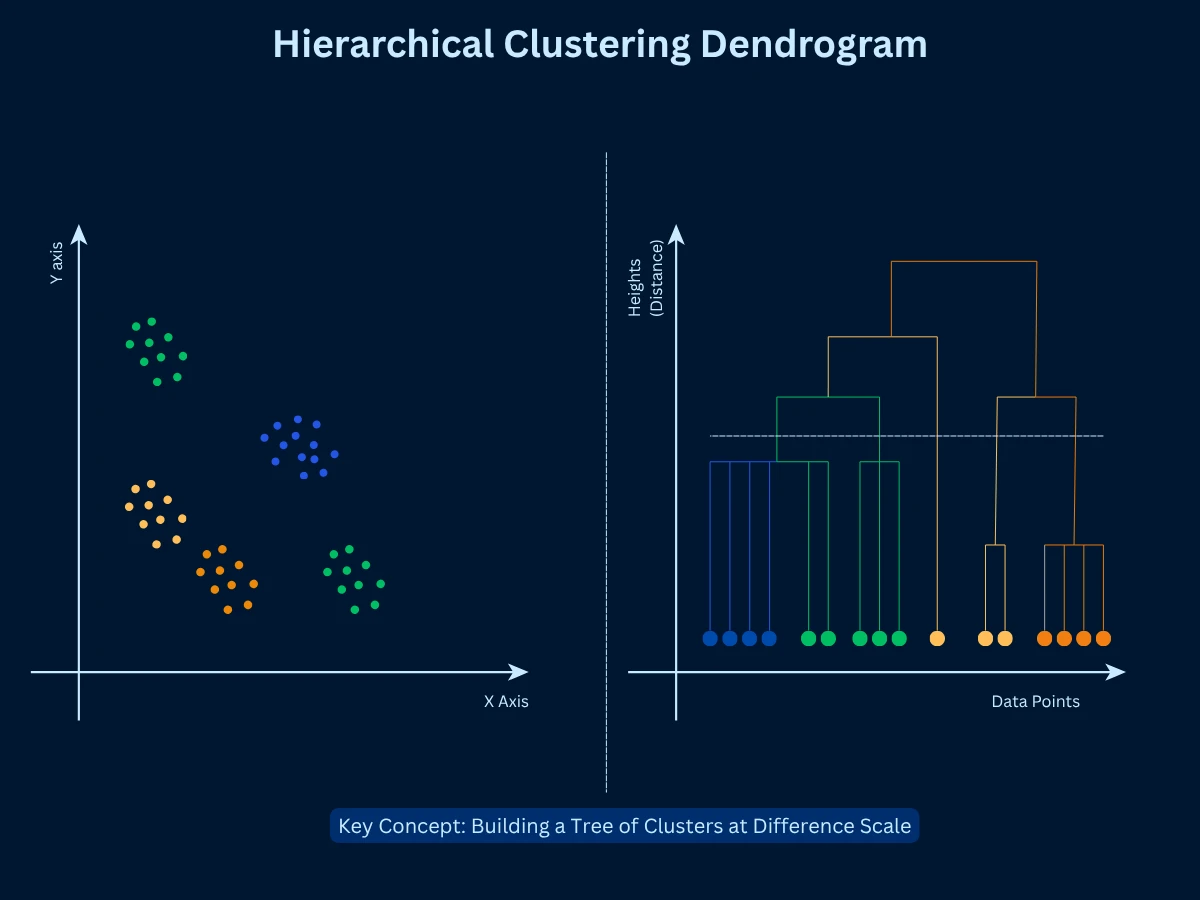
Hierarchical clustering builds a tree (DendrogramDendrogramA tree-shaped diagram that shows how clusters were merged step by step. You can "cut" the tree at any height to get different numbers of clusters from the same result.Example:Like a family tree, but for data - at the top, everyone is one group; at the bottom, each person is their own group.) of nested clusters, allowing you to view data at multiple GranularityGranularityThe level of detail or zoom in your analysis. Fine granularity means many small, specific groups; coarse granularity means fewer, broader groups.Example:Grouping customers by country (coarse) vs by city (fine) - city is higher granularity.. Two approaches exist: AgglomerativeAgglomerativeA bottom-up clustering approach that starts with every single point as its own cluster, then repeatedly merges the two most similar clusters until everything is one group.Example:Like starting with 100 individuals at a party, then grouping people who have things in common, then grouping those groups, until everyone is in one big group. (bottom-up: start with each point as a cluster, merge similar clusters) and divisive (top-down: start with one cluster, split recursively). Agglomerative is more common. The dendrogram shows the merge sequence, and you can cut at any height to get different numbers of clusters.
- How it works: Start with N clusters (one per point), merge the two most similar clusters, repeat until one cluster remains, cut dendrogram at desired height
- Strengths: Don't need to specify K upfront, produces hierarchical structure showing relationships, deterministic results, works with any distance metric
- Limitations: Slow on large datasets (O(n^3) complexity), merge decisions are final (can't undo bad early merges), memory intensive
- Best for: Small to medium datasets, when cluster hierarchy matters, exploratory analysis to determine K, taxonomies
- Common uses: Gene expression analysis, species classification, document organization, social network analysis
1
from sklearn.cluster import AgglomerativeClustering
2
from scipy.cluster.hierarchy import dendrogram, linkage
3
import numpy as np
4
5
# Generate sample data
6
X = np.array([[0, 0], [0, 1], [1, 0],
7
[5, 5], [5, 6], [6, 5],
8
[10, 10], [10, 11], [11, 10]])
9
10
# Perform hierarchical clustering (agglomerative)
11
hierarchical = AgglomerativeClustering(n_clusters=3, linkage='ward')
12
labels = hierarchical.fit_predict(X)
13
14
print(f"Cluster assignments: {labels}")
15
# Output: [0 0 0 1 1 1 2 2 2]
16
17
# Create linkage matrix for dendrogram visualization
18
linkage_matrix = linkage(X, method='ward')
19
20
print(f"\nLinkage matrix (last 5 merges):")
21
print(linkage_matrix[-5:])
22
# Shows which clusters merged, at what distance, forming new clusters
23
24
# Try different numbers of clusters by cutting at different heights
25
for n_clusters in [2, 3, 4]:
26
hc = AgglomerativeClustering(n_clusters=n_clusters, linkage='ward')
27
labels = hc.fit_predict(X)
28
print(f"\nWith {n_clusters} clusters: {labels}")
29
30
# Output:
31
# With 2 clusters: [0 0 0 0 0 0 1 1 1] (merges first two groups)
32
# With 3 clusters: [0 0 0 1 1 1 2 2 2] (optimal - natural groups)
33
# With 4 clusters: [0 0 0 1 2 2 3 3 3] (over-segmentation)
34
35
print(f"\nHierarchical advantage: Can explore multiple K values from one dendrogram")
3. DBSCAN (Density-Based Spatial Clustering)
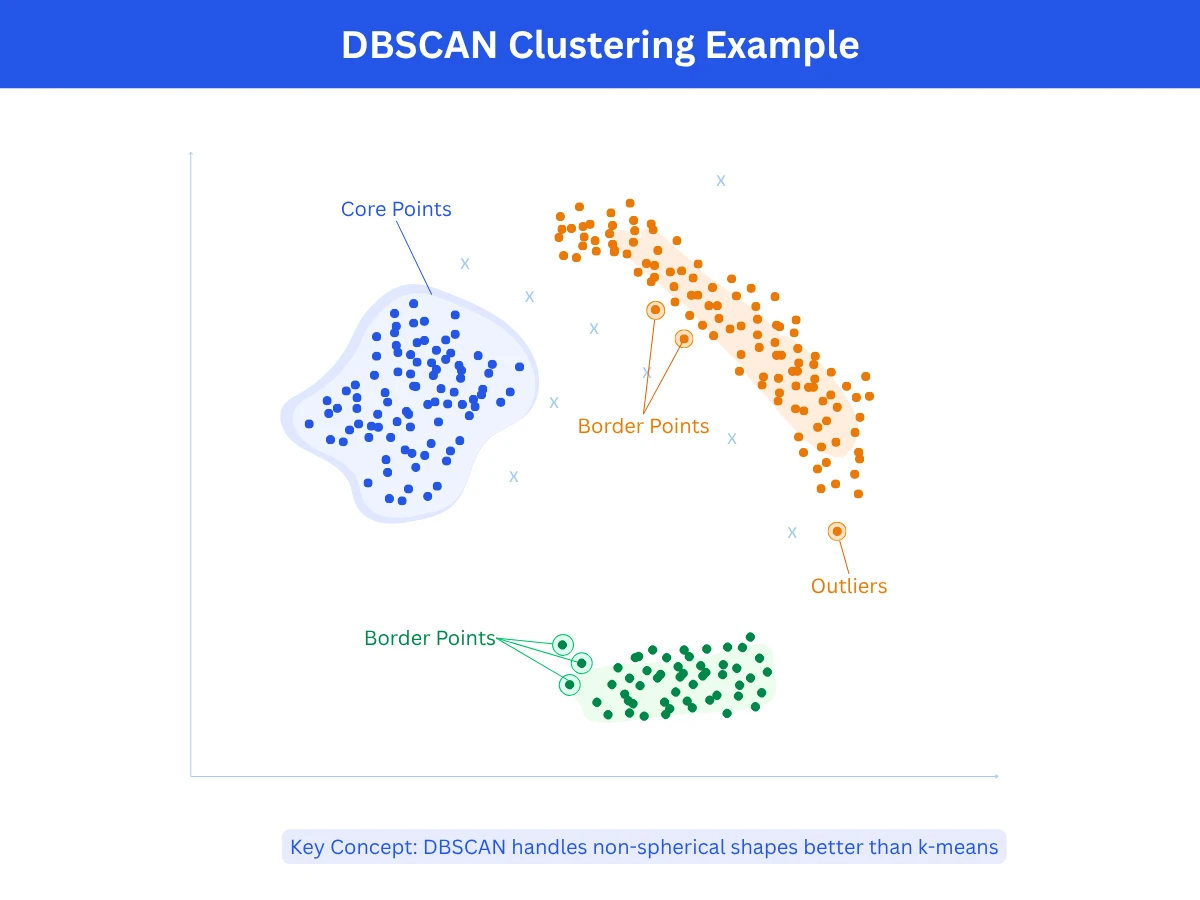
DBSCAN finds clusters by looking for dense areas in your data. Points grouped closely together form a cluster, while isolated points become OutlierOutlierA data point that is significantly different from other observations. Can indicate errors or rare but valid cases.Example:A $50 million mansion in a neighborhood where most houses cost $300K-$500K.. You control two parameters: Epsilon (eps)Epsilon (eps)A distance threshold used in DBSCAN. Any two points closer than epsilon are considered neighbors. Setting it too large merges separate clusters; too small creates too many outliers.Example:Like a social bubble - if epsilon is 2 meters, anyone within 2 meters of you is your neighbor in the cluster. (how close points need to be) and min_points (minimum points required to form a cluster). DBSCAN scans your data, identifies the dense regions, and separates them from sparse areas. It automatically determines the number of clusters, so you don't need to specify it in advance.
- How it works: Start at any point and check how many neighbors it has within epsilon distance. If it has at least min_points neighbors, mark it as a cluster core and add all those neighbors to the cluster. Then repeat the process for each neighbor, expanding the cluster. Points that don't have enough neighbors to form or join a cluster are marked as outliers.
- Strengths: Can discover clusters of any shape (curved, elongated, irregular), automatically identifies outliers without forcing them into clusters, figures out the number of clusters on its own without you specifying K, handles noisy data well by separating signal from noise.
- Limitations: Struggles when clusters have very different densities (one tight cluster and one loose cluster), requires careful tuning of epsilon and min_points parameters which can be tricky to set correctly, runs slower than k-means especially on large datasets, performance degrades in very high-dimensional spaces.
- Best for: Data with irregular or non-spherical cluster shapes, datasets containing noise or outliers that need to be identified separately, spatial or geographic data where clusters naturally form based on proximity, situations where you don't know how many clusters to expect.
- Common uses: Detecting fraudulent transactions or anomalies in financial data, clustering geographic locations like store locations or earthquake epicenters, segmenting objects in images or medical scans, identifying traffic congestion patterns or unusual vehicle movements.

1
from sklearn.cluster import DBSCAN
2
from sklearn.datasets import make_moons
3
import numpy as np
4
5
# Generate crescent-shaped data (impossible for K-Means)
6
X, _ = make_moons(n_samples=200, noise=0.05, random_state=42)
7
8
# Apply DBSCAN clustering
9
dbscan = DBSCAN(eps=0.3, min_samples=5)
10
labels = dbscan.fit_predict(X)
11
12
# Analyze results
13
n_clusters = len(set(labels)) - (1 if -1 in labels else 0)
14
n_outliers = list(labels).count(-1)
15
16
print(f"Number of clusters found: {n_clusters}") # Output: 2
17
print(f"Number of outliers: {n_outliers}") # Output: 4
18
19
# Count points per cluster
20
unique_labels, counts = np.unique(labels[labels != -1], return_counts=True)
21
for label, count in zip(unique_labels, counts):
22
print(f" Cluster {label}: {count} points")
23
# Output:
24
# Cluster 0: 99 points
25
# Cluster 1: 97 points
26
27
# Compare with K-Means (fails on this data)
28
from sklearn.cluster import KMeans
29
kmeans = KMeans(n_clusters=2, random_state=42)
30
kmeans_labels = kmeans.fit_predict(X)
31
32
print(f"\nDBSCAN correctly identifies 2 crescent clusters")
33
print(f"K-Means incorrectly splits crescents vertically")
34
print(f"\nDBSCAN advantage: Discovers arbitrary shapes without knowing K in advance")
4. Gaussian Mixture Models (GMM)
Gaussian Mixture Models assume data comes from a mixture of multiple Gaussian DistributionGaussian DistributionA bell-shaped probability distribution where most values cluster around the center (the average) and taper off symmetrically at the edges. Also called a normal distribution.Example:Human heights follow a Gaussian distribution - most people are average height, fewer are very short or very tall, creating a bell curve. distributions. Instead of hard cluster assignments (point belongs to cluster A), GMM provides Soft AssignmentSoft AssignmentWhen a data point is given a probability of belonging to each cluster rather than being forced into just one. A point might be 70% in cluster A and 30% in cluster B.Example:A music listener who enjoys both jazz and classical - instead of labeling them as one type, soft assignment says they are 60% jazz fan and 40% classical fan. (point has 70% probability of cluster A, 30% of cluster B). This probabilistic approach is more flexible than k-means and can model Elliptical ClustersElliptical ClustersClusters shaped like ovals or elongated blobs, rather than round circles. Gaussian Mixture Models can detect these, unlike K-means which only handles round shapes.Example:Plotting height vs weight for athletes - sprinters might form a tall, narrow oval cluster while weightlifters form a wide, short one. clusters with different sizes and orientations. It uses the Expectation-Maximization (EM)Expectation-Maximization (EM)An iterative algorithm that alternates between two steps: E-step (estimate which cluster each point likely belongs to) and M-step (update the cluster shapes based on those estimates). Repeats until stable.Example:Like sorting laundry by guessing which pile each item belongs to, then updating your sorting rules based on the piles, and repeating until the sorting stops changing. algorithm to find optimal parameters.
- How it works: Start by assuming your data comes from K bell-shaped (Gaussian) distributions. The algorithm alternates between two steps: calculate the probability that each point belongs to each cluster (E-step), then update each cluster's shape and position based on those probabilities (M-step). Repeat this process until the cluster shapes stabilize and stop changing.
- Strengths: Gives you probability scores showing how likely each point belongs to each cluster rather than forcing hard yes/no assignments, can model oval or elliptical cluster shapes with different sizes and orientations, more flexible than k-means because it handles elongated clusters, built on solid statistical theory making results interpretable.
- Limitations: Takes longer to compute than k-means especially on large datasets, assumes your data follows bell-shaped distributions which may not match reality for all datasets, you still need to specify K in advance like k-means, can get stuck in suboptimal solutions depending on initial starting positions.
- Best for: Data that naturally follows bell-shaped patterns when visualized, situations where you need confidence scores or probabilities rather than definite cluster assignments, clusters that are elliptical or oval-shaped rather than circular, modeling data that comes from mixed sources or populations.
- Common uses: Separating foreground objects from backgrounds in images or video processing, identifying different speakers in audio recordings based on voice characteristics, medical imaging to distinguish different tissue types in scans, analyzing gene expression patterns to identify cell types in bioinformatics research.
5. Other Notable Clustering Methods
Several specialized clustering algorithms exist for specific use cases beyond the four main methods above.
- Mean Shift: Imagine each data point gravitating toward the densest crowd nearby. The algorithm shifts each point step-by-step toward areas with the most neighbors until points stop moving. Dense regions where points settle become clusters. You don't need to specify K upfront, and it handles irregular cluster shapes naturally. However, it runs very slowly on datasets with thousands of points. Best for image segmentation (identifying objects in photos) and tracking moving objects in video.
- Spectral Clustering: Treats your data as a network where similar points are connected. It analyzes the mathematical properties of this network to identify natural community structures. Excellent at finding clusters with complex, non-circular shapes that would confuse k-means. However, it's computationally expensive (slow and memory-hungry) and still requires you to specify K. Best used for image segmentation and finding communities in social networks or graphs.
- OPTICS (Ordering Points To Identify the Clustering Structure): An enhanced version of DBSCAN that works better when your data has clusters of different densities. Instead of forcing you to pick one
epsilon value, it creates an ordered list showing how points connect at every density level. This lets you see tight clusters and loose clusters in the same dataset. More flexible than DBSCAN but also more complex to interpret. - Affinity Propagation: Works like a voting system where data points send messages to each other saying 'you should be the cluster center' or 'I should be the cluster center'. After many rounds of message passing, certain points emerge as natural cluster centers (exemplars) and other points join them. Automatically determines the number of clusters without you specifying K. However, it's slow and uses a lot of memory, making it impractical for large datasets (typically limited to a few thousand points).
Choosing the Right Clustering Algorithm
No single clustering algorithm works best for all problems. The right choice depends on your data characteristics, cluster shapes, dataset size, and whether you know the number of clusters in advance.
| Choose This | When | Typical Use Cases |
|---|---|---|
| K-Means | Large dataset, spherical well-separated clusters, K is known | Customer segmentation, image compression, general-purpose clustering |
| Hierarchical | Small dataset, need cluster hierarchy, exploring different K values | Gene expression analysis, taxonomy, understanding data structure |
| DBSCAN | Arbitrary shapes, outliers present, K unknown, spatial data | Anomaly detection, geographic clustering, irregular patterns |
| GMM | Need probability estimates, elliptical clusters, data follows Gaussian distributions | Image segmentation, mixture modeling, soft assignments needed |
| Mean Shift | K unknown, arbitrary shapes, willing to pay computational cost | Image segmentation, object tracking, mode finding |
| Spectral | Non-convex shapes, image segmentation, small-medium datasets | Image segmentation, community detection in graphs |
| Algorithm | Speed | Scalability | Cluster Shapes | Requires K? | Handles Outliers? |
|---|---|---|---|---|---|
| K-Means | Very Fast | Excellent (millions) | Spherical only | Yes | No |
| Hierarchical | Slow | Poor (thousands) | Any shape | No | Somewhat |
| DBSCAN | Medium | Medium (100k) | Arbitrary | No | Yes |
| GMM | Medium | Good (100k+) | Elliptical | Yes | No |
| Mean Shift | Slow | Poor (10k) | Arbitrary | No | Yes |
| Spectral | Slow | Poor (10k) | Non-convex | Yes | Somewhat |
The 80/20 Rule for Clustering
1
from sklearn.cluster import KMeans, DBSCAN, AgglomerativeClustering
2
from sklearn.datasets import make_blobs
3
from sklearn.metrics import silhouette_score
4
import numpy as np
5
6
# Generate test dataset with 3 well-separated clusters
7
X, y_true = make_blobs(n_samples=300, centers=3, cluster_std=0.8, random_state=42)
8
9
# Compare three main clustering algorithms
10
algorithms = [
11
('K-Means', KMeans(n_clusters=3, random_state=42)),
12
('DBSCAN', DBSCAN(eps=1.5, min_samples=5)),
13
('Hierarchical', AgglomerativeClustering(n_clusters=3))
14
]
15
16
print("Algorithm Comparison on Same Dataset")
17
print("=" * 70)
18
19
for name, algorithm in algorithms:
20
# Fit and predict
21
if name == 'DBSCAN':
22
labels = algorithm.fit_predict(X)
23
n_clusters = len(set(labels)) - (1 if -1 in labels else 0)
24
n_outliers = list(labels).count(-1)
25
else:
26
labels = algorithm.fit_predict(X)
27
n_clusters = len(set(labels))
28
n_outliers = 0
29
30
# Calculate silhouette score (skip if too few clusters)
31
if n_clusters > 1 and n_clusters < len(X) - 1:
32
silhouette = silhouette_score(X, labels)
33
else:
34
silhouette = 0.0
35
36
print(f"\n{name}:")
37
print(f" Clusters found: {n_clusters}")
38
print(f" Outliers: {n_outliers}")
39
print(f" Silhouette score: {silhouette:.3f}")
40
41
# Points per cluster
42
unique_labels = [l for l in set(labels) if l != -1]
43
for label in sorted(unique_labels):
44
count = list(labels).count(label)
45
print(f" Cluster {label}: {count} points")
46
47
# Output:
48
# K-Means:
49
# Clusters found: 3
50
# Outliers: 0
51
# Silhouette score: 0.632
52
# Cluster 0: 100 points
53
# Cluster 1: 100 points
54
# Cluster 2: 100 points
55
#
56
# DBSCAN:
57
# Clusters found: 3
58
# Outliers: 2
59
# Silhouette score: 0.645
60
# Cluster 0: 98 points
61
# Cluster 1: 100 points
62
# Cluster 2: 100 points
63
#
64
# Hierarchical:
65
# Clusters found: 3
66
# Outliers: 0
67
# Silhouette score: 0.632
68
# Cluster 0: 100 points
69
# Cluster 1: 100 points
70
# Cluster 2: 100 points
71
72
print(f"\nConclusion: All 3 algorithms perform similarly on well-separated spherical clusters")
73
print(f"DBSCAN slightly edges out with 0.645 silhouette and identifies 2 outliers")
Evaluating Clustering Quality
Unlike supervised learning, clustering has no ground truth labels to measure accuracy against. How do you know if your clustering is good? Several metrics and techniques exist for evaluating cluster quality without labels.
Internal Validation Metrics
Internal metrics evaluate clustering quality using only the data and cluster assignments - no external information needed. They measure how compact clusters are (points within a cluster are similar) and how separated clusters are (different clusters are distinct).
- Silhouette Score (-1 to 1): Measures how similar each point is to its own cluster compared to other clusters. Values near 1 indicate excellent clustering, near 0 indicates overlapping clusters, negative values indicate misassignment.
- Davies-Bouldin Index: Ratio of within-cluster distances to between-cluster distances. Lower is better. Measures cluster compactness vs separation.
- Calinski-Harabasz Index: Ratio of between-cluster dispersion to within-cluster dispersion. Higher is better. Rewards compact, well-separated clusters.
- Inertia (Within-Cluster Sum of Squares): Sum of squared distances from points to their cluster centroids. Lower is better, but decreases as K increases (can overfit).
Choosing the Optimal Number of Clusters (K)
For algorithms requiring K (k-means, GMM), choosing the right number of clusters is critical. Several techniques help determine optimal K.
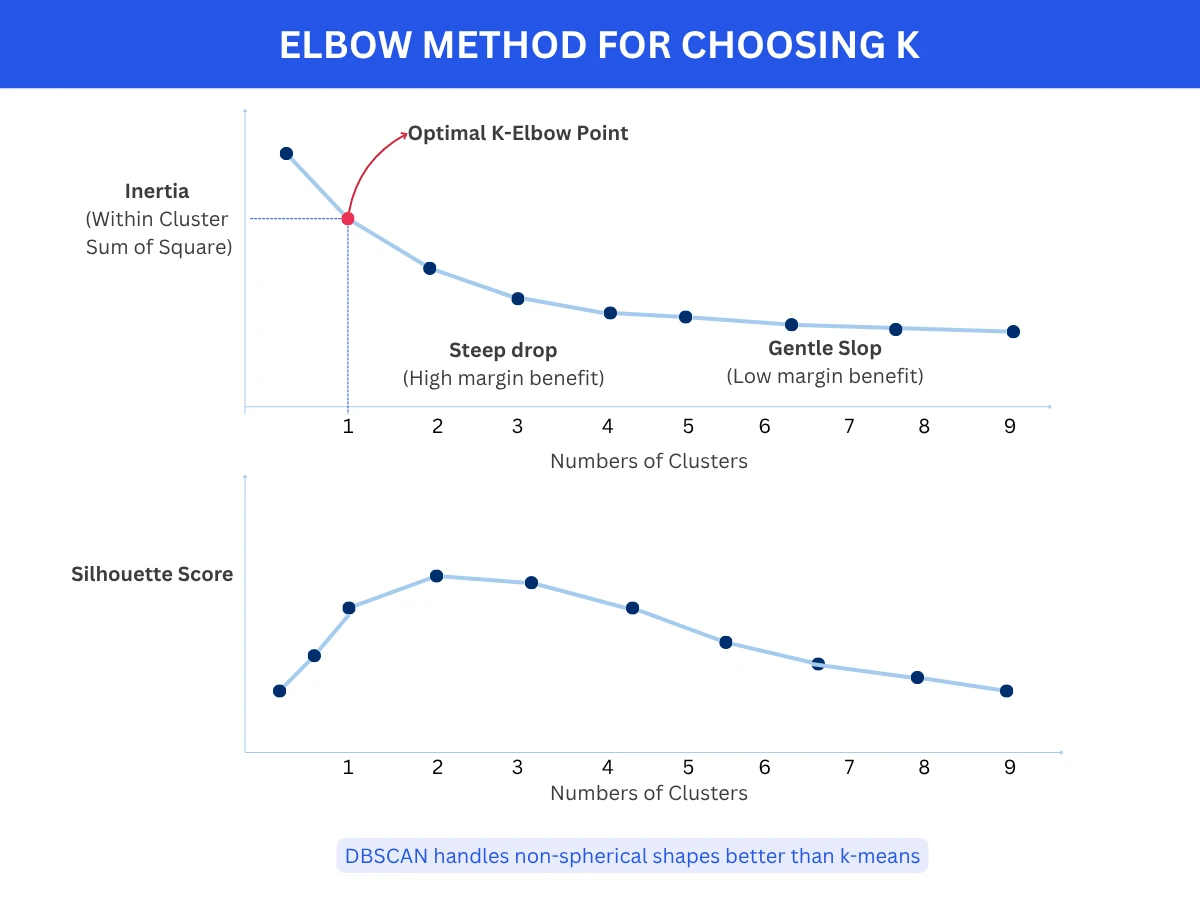
- Elbow Method: Plot inertia vs K. Look for the 'elbow' where adding more clusters provides diminishing returns. The elbow point suggests optimal K.
- Silhouette Analysis: Plot silhouette score vs K. Choose K with highest average silhouette score indicating best-defined clusters.
- Gap Statistic: Compare inertia to null reference distribution. Choose K where gap is largest, indicating clustering structure significantly better than random.
- Domain Knowledge: Sometimes business requirements dictate K (e.g., divide customers into exactly 4 tiers: budget, standard, premium, luxury).
1
from sklearn.cluster import KMeans
2
from sklearn.datasets import make_blobs
3
from sklearn.metrics import silhouette_score
4
import numpy as np
5
6
# Generate data with 4 natural clusters
7
X, _ = make_blobs(n_samples=400, centers=4, cluster_std=0.6, random_state=42)
8
9
# Elbow Method: Try different K values
10
inertias = []
11
silhouette_scores = []
12
K_range = range(2, 11)
13
14
for k in K_range:
15
kmeans = KMeans(n_clusters=k, random_state=42, n_init=10)
16
kmeans.fit(X)
17
inertias.append(kmeans.inertia_)
18
silhouette_scores.append(silhouette_score(X, kmeans.labels_))
19
20
print("K-value evaluation:")
21
print("K | Inertia | Silhouette | Assessment")
22
print("-" * 50)
23
for k, inertia, sil in zip(K_range, inertias, silhouette_scores):
24
assessment = ""
25
if k == 4:
26
assessment = "<-- OPTIMAL (elbow + high silhouette)"
27
print(f"{k:2d} | {inertia:8.1f} | {sil:10.3f} | {assessment}")
28
29
# Output:
30
# 2 | 16250.3 | 0.532 |
31
# 3 | 7854.2 | 0.618 |
32
# 4 | 4523.1 | 0.651 | <-- OPTIMAL (elbow + high silhouette)
33
# 5 | 3612.8 | 0.612 |
34
# 6 | 2985.4 | 0.589 |
35
# ...
36
37
print(f"\nRecommendation: K=4")
38
print(f"- Inertia drops steeply up to K=4, then flattens (elbow)")
39
print(f"- Silhouette score peaks at K=4 (0.651)")
40
print(f"- K=4 matches true number of clusters in data")
No Perfect Clustering
Real-World Applications
Clustering powers applications across every industry, from customer segmentation to medical diagnosis to astronomy. Understanding these use cases helps you recognize clustering opportunities in your own work.
| Domain | Application | Algorithm Used | Business Impact |
|---|---|---|---|
| E-commerce | Customer segmentation by purchase behavior | K-Means, GMM | Personalized marketing increased conversion 20-30% |
| Healthcare | Identifying patient subgroups from genetic data | Hierarchical, GMM | Discovered new cancer subtypes enabling targeted treatments |
| Marketing | Market segmentation and persona development | K-Means | Reduced ad spend waste by 40% through targeted campaigns |
| Image Processing | Image segmentation and compression | K-Means, Mean Shift | Reduced image file sizes by 70% while maintaining quality |
| Fraud Detection | Detecting unusual transaction patterns | DBSCAN, Isolation Forest | Identified fraud patterns 2-3x faster than rule-based systems |
| Astronomy | Classifying celestial objects from telescope data | DBSCAN, Hierarchical | Discovered previously unknown galaxy types |
| Biology | Gene expression pattern discovery | Hierarchical, K-Means | Identified disease biomarkers and drug targets |
| Social Networks | Community detection in user networks | Spectral, Hierarchical | Improved content recommendations by 25% |
| Document Management | Organizing documents by topic | K-Means, LDA | Reduced search time by 60% through automatic categorization |
| Telecommunications | Network traffic pattern analysis | DBSCAN, K-Means | Detected network anomalies 15 minutes faster |
How Spotify Discovers Music Genres
Frequently Asked Questions
01 How do I know if clustering will work on my data?
Visualize your data first! If your data has 2-3 dimensions, plot it and look for natural groupings. If high-dimensional, use dimensionality reduction (PCA, t-SNE) to project to 2D for visualization. If you see clear groupings, clustering will likely work well. If data looks uniformly distributed with no patterns, clustering may not be meaningful. Also check: Do similar data points make domain sense grouped together? If yes, proceed with clustering.
02 Should I standardize features before clustering?
YES, almost always! Distance-based algorithms (k-means, hierarchical, DBSCAN) are sensitive to feature scales. If one feature ranges 0-1 while another ranges 0-10000, the large-scale feature dominates distance calculations. SOLUTION: Standardize features to mean=0, std=1 using StandardScaler, or normalize to 0-1 range using MinMaxScaler. Exception: if all features already have similar scales and meaningful units, standardization may not be necessary.
03 Why does k-means give different results each time I run it?
K-means initializes centroids randomly, and different starting positions can lead to different local optima. SOLUTIONS: (1) Run k-means multiple times (10-20 runs) with different random seeds and pick the result with lowest inertia, (2) Use k-means++ initialization (default in most libraries) which chooses better starting centroids, (3) Set random_state parameter for reproducibility during development, (4) Consider if results are fundamentally different or just cluster labels are permuted (clusters [A,B,C] vs [C,A,B] are the same).
04 Can clustering handle categorical features?
Standard distance-based algorithms (k-means, DBSCAN) expect numerical features. For categorical features: (1) Use one-hot encoding to convert categories to binary features, (2) Use specialized algorithms like k-modes (like k-means but for categorical data) or k-prototypes (handles mixed numerical and categorical), (3) For hierarchical clustering, use appropriate distance metrics like Hamming distance for categorical data, (4) Encode ordinal categories as integers if ordering is meaningful (small, medium, large - 1, 2, 3).
05 How do I interpret and name clusters after clustering?
After clustering, analyze each cluster's characteristics: (1) Calculate mean/median of each feature per cluster to understand cluster centers, (2) Identify which features most distinguish each cluster (high variance between clusters, low within), (3) Examine representative examples from each cluster, (4) Use domain knowledge to assign meaningful names, (5) Visualize clusters using parallel coordinates or radar charts showing feature profiles, (6) Validate clusters make business sense and are actionable.
06 What's the difference between k-means and k-medoids?
K-means: Uses cluster centers (centroids) calculated as the mean of all points in the cluster. Centroid may not be an actual data point. Fast but sensitive to outliers. K-medoids (PAM): Uses actual data points as cluster centers (medoids). More robust to outliers since medoid must be a real point. Slower and more computationally expensive. Use k-medoids when: (1) outliers are present, (2) you need cluster centers to be actual data points, (3) you have a custom distance metric.
07 Can I use clustering for dimensionality reduction?
Yes! Clustering can be used for feature engineering: (1) Cluster your data, (2) Use cluster membership as a new categorical feature, (3) Use distance to each cluster centroid as new features - this creates K new features from original features. For example, cluster customers into 5 segments, then use segment membership (0-4) as a feature for supervised learning. However, dedicated dimensionality reduction methods (PCA, t-SNE, UMAP) are usually better for pure dimensionality reduction.
08 How do I cluster high-dimensional data effectively?
High-dimensional data suffers from the curse of dimensionality - distances become less meaningful as dimensions increase. SOLUTIONS: (1) Apply dimensionality reduction first (PCA to reduce to 10-50 dimensions) then cluster, (2) Use feature selection to keep only most relevant features, (3) Consider specialized algorithms like spectral clustering that work better in high dimensions, (4) Use cosine similarity instead of Euclidean distance for sparse data, (5) For very high dimensions (> 1000), use deep learning approaches like autoencoders to learn low-dimensional representations first.
Ready to Dive Deeper?
You now understand the most important clustering algorithms, when to use each one, and how to evaluate clustering quality. Your next step is exploring dimensionality reduction techniques or diving deeper into specific clustering algorithms.
Your Next Step
Clustering is both an art and a science. The algorithms are mathematical and deterministic, but interpreting results requires domain expertise and judgment. I've seen teams run perfect clustering algorithms and get meaningless results because they didn't understand their data. The best clustering projects start with: What question am I trying to answer? What would make a cluster useful? Then choose the algorithm that answers those questions, not the fanciest one.
